
In a recent study published in the journal Infectious Diseases of Poverty, researchers used transmission-locality data for Oropouche virus (OROV) and high-resolution vegetation phenology from satellite data to develop spatial epidemiology models to understand and predict the spread of the zoonotic vector-borne disease Oropouche fever.
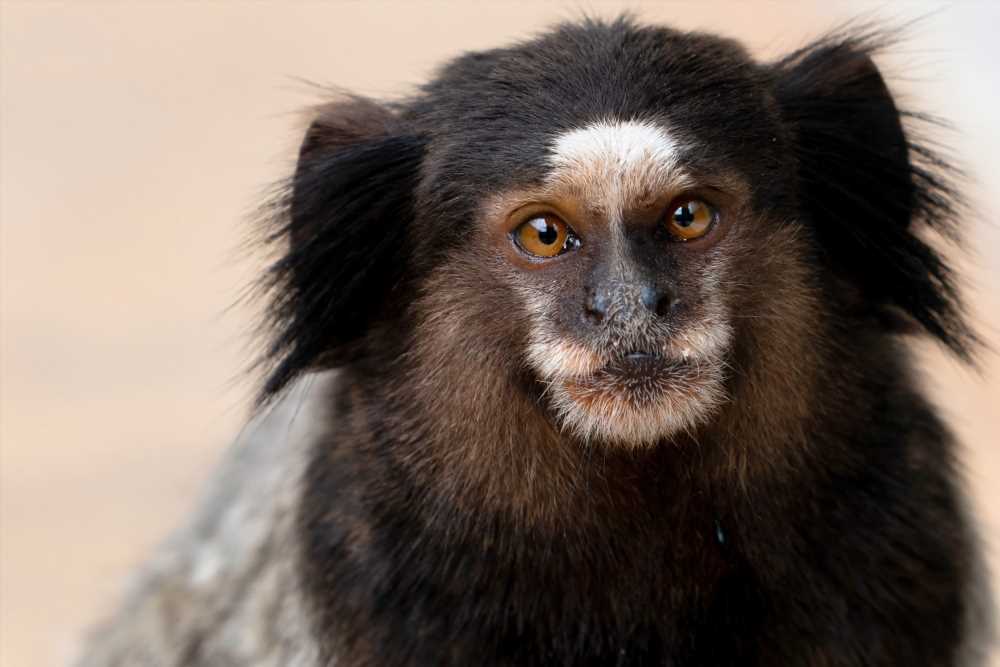
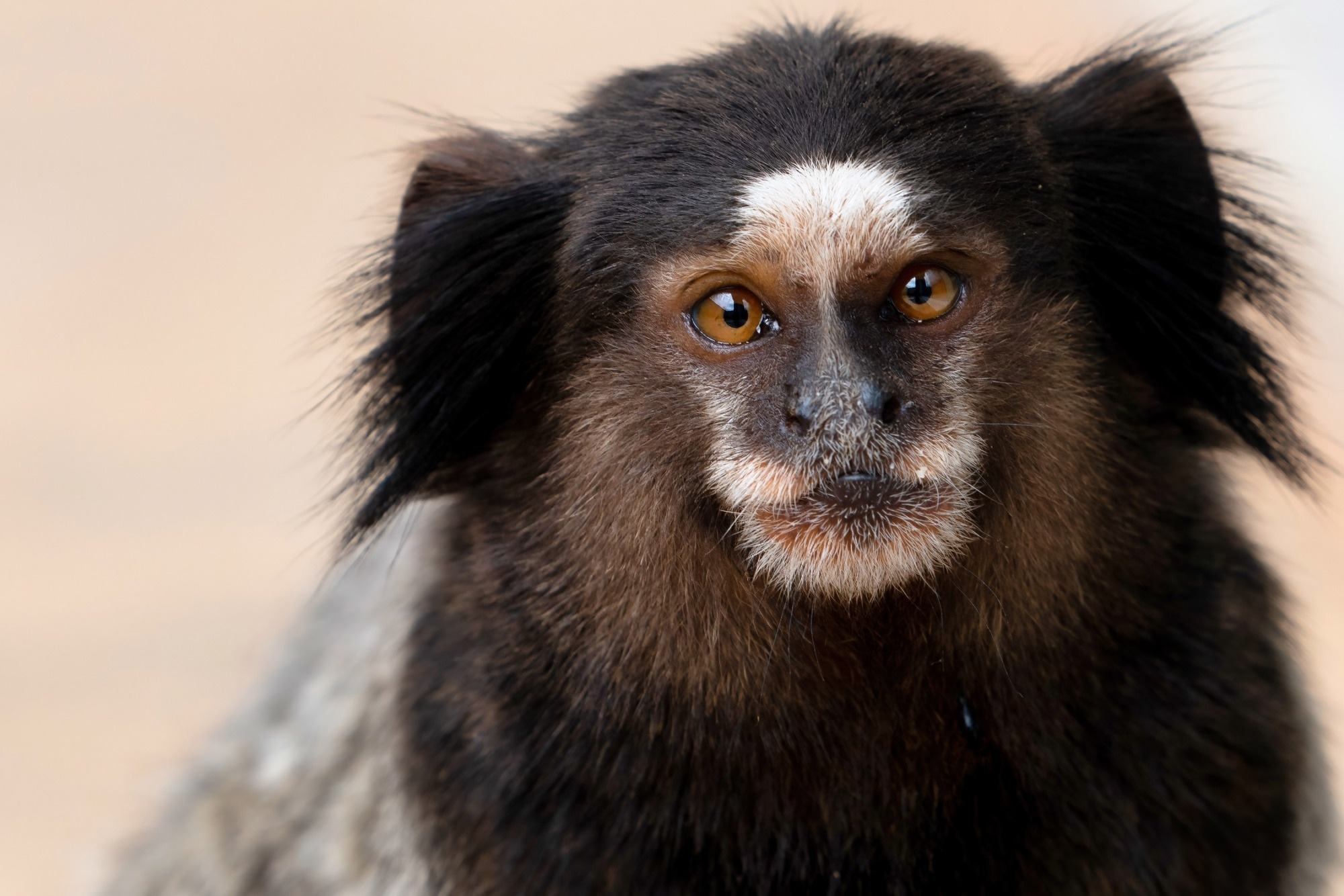 Study: Transmission risk of Oropouche fever across the Americas. Image Credit: pauloalberto82 / Shutterstock
Study: Transmission risk of Oropouche fever across the Americas. Image Credit: pauloalberto82 / Shutterstock
Background
Approximately 17% of infectious diseases worldwide and a loss of about 52,000 disability-adjusted life years are attributed to vector-borne diseases. Although vector-borne diseases are more common in the tropics, the incidence of vector-borne diseases in other regions is increasing due to globalization, large-scale landscape changes, and climate change. Furthermore, a lack of understanding of the epidemiological and ecological factors that drive the transmission of these viruses and the emergence of novel pathogens is challenging the existing public health systems.
The Oropouche virus belongs to the genus Orthobunyavirus and was first described in 1954 in Trinidad and Tobago. The symptoms of Oropouche fever include fever, myalgia, and headaches, which are also common with other vector-borne diseases such as Zika, dengue, and Mayaro fevers. The Oropouche virus is a negative sense ribonucleic acid (RNA) virus whose sylvatic cycle is maintained in wildlife hosts such as non-human primates and sloths and is transmitted through arthropod vectors such as Culex mosquitoes and midges. Given that OROV has infected at least 500,000 individuals in Latin American regions and has the potential to trigger an epidemic, it is essential to understand the factors that influence the spread of the virus.
About the study
In the present study, the researchers used a framework of biogeographic risk mapping using data on human cases of Oropouche fever and landscape data derived from satellite images. A range of modeling protocols was tested to identify the approaches that provide the best predictive capabilities and robust descriptions. These models were then used to identify potential areas where Oropouche fever could occur and areas that had unknown febrile syndromes that could potentially be caused by OROV.
Furthermore, the researchers also studied how changes in the landscape can impact the emergence of OROV and used this understanding to estimate the number of people that might be at risk. Occurrence data for Oropouche fever was compiled using published studies and reports on human cases.
Climatic predictors were obtained at a resolution of about 7 km from a repository for satellite-derived data on humidity and temperature. The three predictors — annual mean temperature, annual mean specific humidity, and annual temperature range — are important factors determining the distribution of midges (Culicoides parensis) and other dipteran vectors. Furthermore, the dispersal capacity of the vector and wildlife host species is another important parameter since it determines the ecological niche and species distribution models.
A hypervolume approach was used to integrate the data, and the selected parameters were used to calibrate and evaluate the models. The selected models were then used to examine how vegetation cover differed between areas of OROV outbreaks and other random locations. Finally, the overlaps between OROV risk maps and human population distributions were used to estimate the proportion of humans at risk of Oropouche fever.
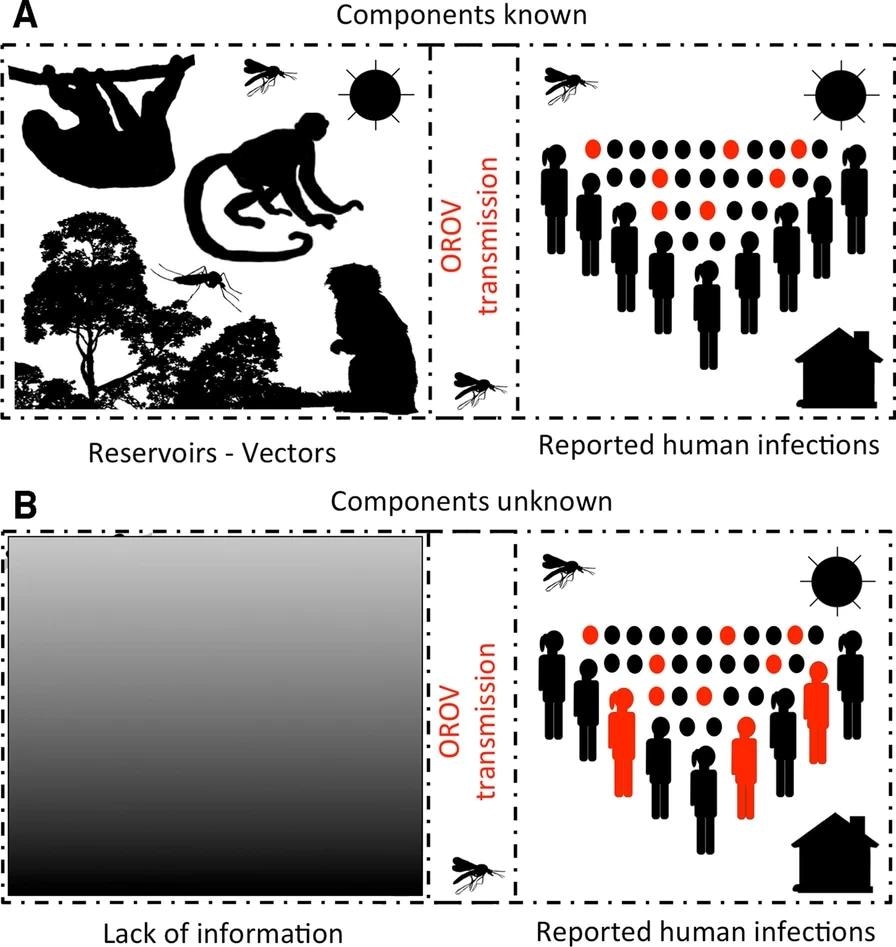
Schematic representation of component or black box-based strategies for infectious disease species distribution modeling. In well-known systems, disease models should aim to model each component driving the life cycle of the pathogen to better characterize its distribution (A). However, for Oropouche virus (OROV), there are multiple gaps in knowledge to actually make assumptions about its sylvatic cycle, specifically, reservoirs and vectors driving epizootics are poorly represented in the scientific literature (B). For these cases, the presence of human outbreaks allows a black box modeling where we assume that detected human cases represent the manifestation of the entire virus cycle despite the unknowns surrounding its components. Silhouettes developed with Adobe Photoshop Elements
Results
The results reported that the selected models predicted that the tropical regions of Latin America continued to be risk areas for OROV transmission, despite the inclusion of various environmental predictors and different study areas. The estimates also reported that close to five million individuals were at risk of OROV exposure. Furthermore, landscape changes associated with vegetation loss were linked to the Oropouche fever outbreak risk.
However, the researchers caution that there is a degree of uncertainty associated with these projections due to the limited data on the epidemiology of the virus. There have been examples of outbreaks in regions that have climatic conditions outside of the range where most transmissions occur.
The findings also showed that a hypervolume modeling approach can be used to understand the geographic and ecological patterns in disease occurrence and predict transmission risk. Vegetation loss seems to be one of the main drivers of Oropouche fever occurrence.
Conclusions
To summarize, the study used a hypervolume epidemiological modeling approach to estimate the transmission risk and predict potential areas of occurrence of Oropouche fever. The tropical regions of Latin America are at a high risk of future outbreaks, with two to five million people at risk of OROV exposure. Furthermore, vegetation loss was seen to be associated with an increased risk of Oropouche fever.
- Romero-Alvarez, D., Escobar, L. E., Auguste, A. J., Del Valle, S. Y., & Manore, C. A. (2023). Transmission risk of Oropouche fever across the Americas. Infectious Diseases of Poverty, 12(1). https://doi.org/10.1186/s40249-023-01091-2, https://www.frontiersin.org/articles/10.3389/fped.2023.1130179/full
Posted in: Medical Science News | Medical Research News | Disease/Infection News
Tags: Climate Change, Disability, Epidemiology, Fever, Health Systems, Infectious Diseases, Pathogen, Poverty, Public Health, Ribonucleic Acid, RNA, Virus
.jpg)
Written by
Dr. Chinta Sidharthan
Chinta Sidharthan is a writer based in Bangalore, India. Her academic background is in evolutionary biology and genetics, and she has extensive experience in scientific research, teaching, science writing, and herpetology. Chinta holds a Ph.D. in evolutionary biology from the Indian Institute of Science and is passionate about science education, writing, animals, wildlife, and conservation. For her doctoral research, she explored the origins and diversification of blindsnakes in India, as a part of which she did extensive fieldwork in the jungles of southern India. She has received the Canadian Governor General’s bronze medal and Bangalore University gold medal for academic excellence and published her research in high-impact journals.
Source: Read Full Article